PCRISPomyces-2: Difference between revisions
From ActinoBase
No edit summary |
No edit summary |
||
| Line 3: | Line 3: | ||
[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4459934/ The paper describing pCRISPomyces2] | [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4459934/ The paper describing pCRISPomyces2] | ||
The protocol from Matt Hutchings lab - [http://www.hutchingslab.uk/downloads/pCRISPOmyces2.pdf | The protocol from Matt Hutchings lab - [http://www.hutchingslab.uk/downloads/pCRISPOmyces2.pdf download as a PDF]. | ||
The plasmid is available from AddGene under MTA: [https://www.addgene.org/61737/ click here] | The plasmid is available from AddGene under MTA: [https://www.addgene.org/61737/ click here] | ||
[[file:CRISPR.png]] | [[file:CRISPR.png]] | ||
Revision as of 20:16, 18 July 2019
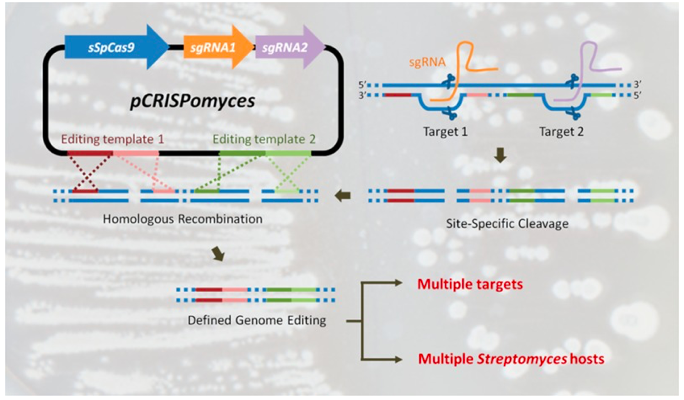
This plasmid can be used for precise CRISPR/Cas9-mediated genome engineering of Streptomyces strains. To our knowledge it has been successfully used in S. albidoflavus, S. coelicolor, S. formicae, S. lividans and S. venezuelae. Mutations have ranged from 1-100kbp deletions, precise codon changes to alter amino acids and insertions to add Flag-tags to proteins encoded at their native loci.
The paper describing pCRISPomyces2
The protocol from Matt Hutchings lab - download as a PDF.
The plasmid is available from AddGene under MTA: click here