PCRISPomyces-2: Difference between revisions
From ActinoBase
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
This plasmid can be used for precise CRISPR/Cas9-mediated genome engineering of <em>Streptomyces</em> strains. To our knowledge it has been successfully used in [https://doi.org/10.1128/mSphere.00305-16 S. albidoflavus], S. coelicolor, [https://pubs.rsc.org/en/content/articlelanding/2017/SC/C6SC04265A#!divAbstract S. formicae] and S. venezuelae. | This plasmid can be used for precise CRISPR/Cas9-mediated genome engineering of <em>Streptomyces</em> strains. To our knowledge it has been successfully used in [https://doi.org/10.1128/mSphere.00305-16 S. albidoflavus], S. coelicolor, [https://pubs.rsc.org/en/content/articlelanding/2017/SC/C6SC04265A#!divAbstract S. formicae], [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4459934/ S. lividans] and S. venezuelae. | ||
Revision as of 19:34, 17 July 2019
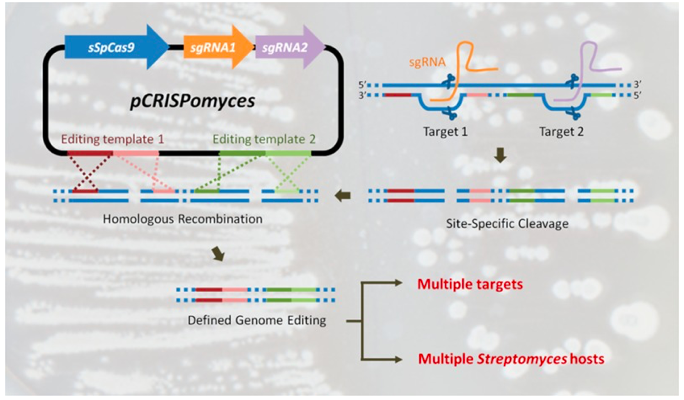
This plasmid can be used for precise CRISPR/Cas9-mediated genome engineering of Streptomyces strains. To our knowledge it has been successfully used in S. albidoflavus, S. coelicolor, S. formicae, S. lividans and S. venezuelae.
The paper describing pCRISPomyces2
The plasmid is available from AddGene under MTA: click here