Streptomyces formicae: Difference between revisions
No edit summary |
No edit summary |
||
| Line 7: | Line 7: | ||
<strong>Further reading:</strong> | <strong>Further reading:</strong> | ||
Qin Z, Devine R, Wilkinson KA, Hutchings MI and Wilkinson B (2019). Aromatic polyketide biosynthesis: fidelity, evolution and engineering. Nature Comms, accepted [https://nature-research-under-consideration.nature.com/users/37265-nature-communications/posts/45836-aromatic-polyketide-biosynthesis-fidelity-evolution-and-engineering Abstract]. | Qin Z, Devine R, Wilkinson KA, Hutchings MI<sup>*</sup> and Wilkinson B<sup>*</sup> (2019). Aromatic polyketide biosynthesis: fidelity, evolution and engineering. Nature Comms, accepted [https://nature-research-under-consideration.nature.com/users/37265-nature-communications/posts/45836-aromatic-polyketide-biosynthesis-fidelity-evolution-and-engineering Abstract]. | ||
Holmes NA, Devine R, Qin Z, Seipke RF, Wilkinson B, Hutchings MI (2018). Complete genome sequence of Streptomyces formicae KY5, the formicamycin producer. J Biotechnol. 265:116-118. [https://www.ncbi.nlm.nih.gov/pubmed/29191667 Abstract] | Holmes NA, Devine R, Qin Z, Seipke RF, Wilkinson B<sup>*</sup>, Hutchings MI<sup>*</sup> (2018). Complete genome sequence of Streptomyces formicae KY5, the formicamycin producer. J Biotechnol. 265:116-118. [https://www.ncbi.nlm.nih.gov/pubmed/29191667 Abstract] | ||
Qin Z, Munnoch JT, Devine R, Holmes NA, Seipke RF, Wilkinson KA, Wilkinson B, Hutchings MI (2017). Formicamycins, antibacterial polyketides produced by Streptomyces formicae isolated from African Tetraponera plant-ants. Chem Sci. 8:3218-3227. [https://www.ncbi.nlm.nih.gov/pubmed/28507698 Abstract] | Qin Z, Munnoch JT, Devine R, Holmes NA, Seipke RF, Wilkinson KA, Wilkinson B<sup>*</sup>, Hutchings MI<sup>*</sup> (2017). Formicamycins, antibacterial polyketides produced by Streptomyces formicae isolated from African Tetraponera plant-ants. Chem Sci. 8:3218-3227. [https://www.ncbi.nlm.nih.gov/pubmed/28507698 Abstract] | ||
Revision as of 09:36, 3 July 2019
This new species of Streptomyces was isolated in 2013 from the domatia (nest) of the Kenyan plant ant Tetraponera penzigi. It contains at least 45 secondary metabolite biosynthetic gene clusters, the vast majority of which are predicted to encode novel products. It also makes the formicamycins, which are potent against MRSA and have a high natural barrier to resistance.
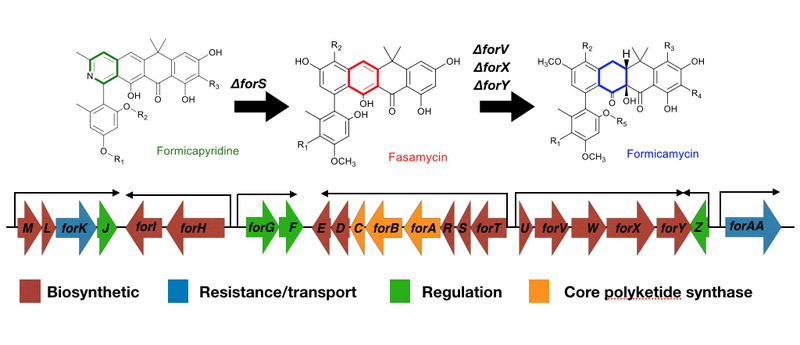
Figure 1. Streptomyces formicae is best known for producing formicamycins. The for biosynthetic gene cluster (BGC) encodes the biosynthesis of three groups of polyketides, the fasamycins, formicamycins and formicapyridines.
Further reading:
Qin Z, Devine R, Wilkinson KA, Hutchings MI* and Wilkinson B* (2019). Aromatic polyketide biosynthesis: fidelity, evolution and engineering. Nature Comms, accepted Abstract.
Holmes NA, Devine R, Qin Z, Seipke RF, Wilkinson B*, Hutchings MI* (2018). Complete genome sequence of Streptomyces formicae KY5, the formicamycin producer. J Biotechnol. 265:116-118. Abstract
Qin Z, Munnoch JT, Devine R, Holmes NA, Seipke RF, Wilkinson KA, Wilkinson B*, Hutchings MI* (2017). Formicamycins, antibacterial polyketides produced by Streptomyces formicae isolated from African Tetraponera plant-ants. Chem Sci. 8:3218-3227. Abstract