PIJ10257: Difference between revisions
From ActinoBase
No edit summary |
(→Use) |
||
| Line 4: | Line 4: | ||
==Use== | ==Use== | ||
This vector (originally published by the Buttner group <sup>1</sup>) can be used to introduce a gene or set of genes | This vector (originally produced and published by the Buttner group <sup>1</sup>) can be used to introduce a gene or set of genes into <em> Streptomyces </em> under the control of the constitutive <em> ermE*</em> promoter <sup>2</sup>. This can be useful for over expression or complementation experiments. | ||
==Features== | ==Features== | ||
Revision as of 13:48, 25 October 2019
Use
This vector (originally produced and published by the Buttner group 1) can be used to introduce a gene or set of genes into Streptomyces under the control of the constitutive ermE* promoter 2. This can be useful for over expression or complementation experiments.
Features
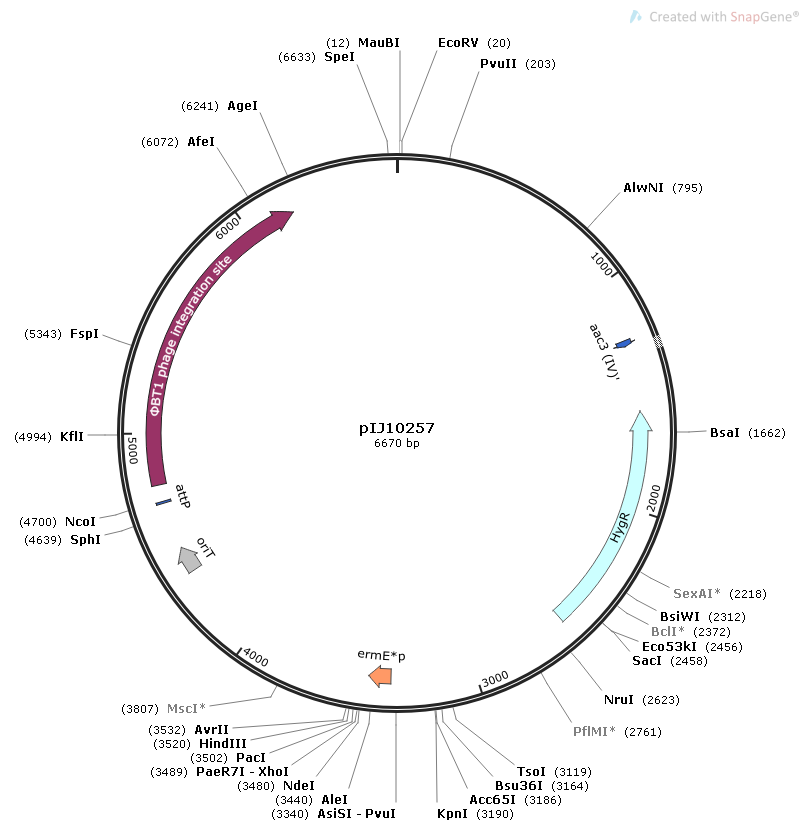
- Hygromycin resistance gene
- ermE* promoter2
- ΦBT1 phage integration site
- Origin of transfer oriT from RK2
History
Backbone vector pMS81 was modified by cloning in a 330 bp fragment containing ermE*, a ribosome binding site and a multicloning site isolated from pIJ8723, to create pIJ102571.
Map
Sequence links
References
- Hong, H., Hutchings, M. I., Hill, L. M., Buttner, M. J. (2005). The Role of the Novel Fem Protein VanK in Vancomycin Resistance in Streptomyces coelicolor. Journal of Biological Chemistry, 280, pp. 13055-13061. doi: 10.1074/jbc.M413801200
- Bibb, M. J., Janssen, G. R., and Ward, J. M. (1985). Cloning and analysis of the promoter region of the erythromycin resistance gene (ermE) of Streptomyces erythraeus. ‘’Gene’’, 38, pp. 215-226. doi: 10.1016/0378-1119(85)90220-3